
Here's the command I used for downloading: ascp -QT -l 300m -i ~/.aspera/connect/etc/asperaweb_id_dsa.openssh. The estimated completion time via standard FTP was 12 hours for about 32GB of data ascp has reduced that estimated download time to about an hour. I am using the version called Ubuntu Linux 64 bit architecture - non-sudo tar archive At the time of writing it is 2.10. In my case, I've just started downloading some files from a MinION sequencing run. Head to the SRA Toolkit GitHub download page and copy the link for the latest version of the toolkit. Your command should look similar to this on Unix: ascp -QT -l 300m -i /etc/asperaweb_id_dsa.openssh On a related note, just in case someone sees this and EBI/ENA is an option, there's a great guide for how to do file transfer using Aspera on the EBI web site: 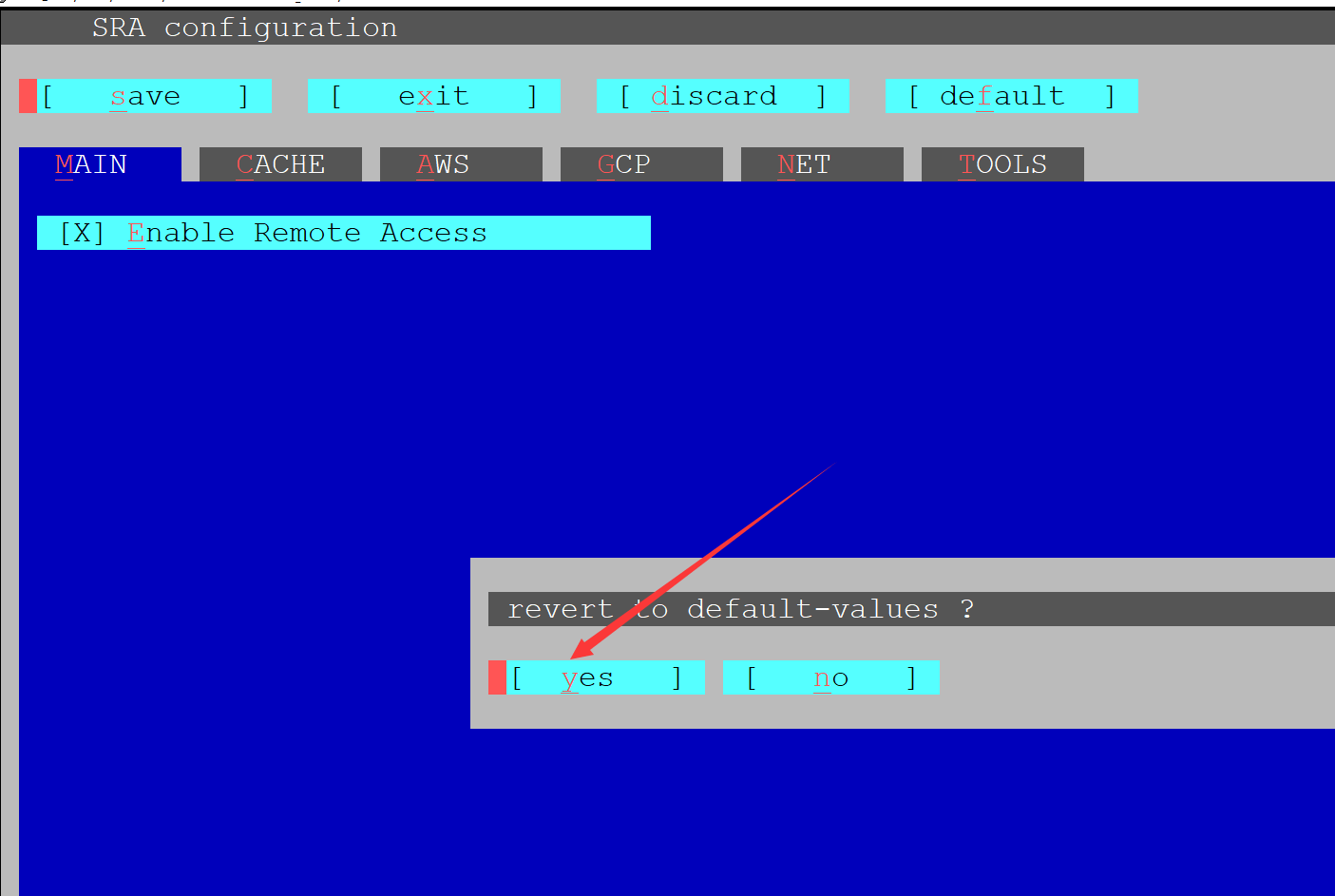
NCBI also has an online book about best practises for downloading data from their servers. This is unlikely to help if bandwidth is being throttled, but it might help if there's a bit of congestion through the regular methods: There's a chance that a home network might be faster, but you're likely to get the fastest connection to NCBI by using an academic system that is linked to NCBI via a research network.Īnother possibility is using Aspera for downloads. AWS may be deliberately throttling the Internet connection to limit the likelihood that people will use it for undesirable things. Proximity to NCBI may not necessarily give you the fastest transfer speed. Is there a better way to fetch this data that wouldn't take multiple days? Surely the SRA must be capable of moving data at higher rates that this? I've also looked, and unfortunately the datasets I'm interested have not been mirrored out to ENA or the Japanese archive, so it looks like I'm stuck working with the SRA. They do appear to still be running - the fastq files continue to grow and their timestamps continue to update.Īt this rate it's going to take me days or weeks just to download the datasets. In 18 hours they've downloaded a total of 33GB of data. They're running on a large AWS instance in the US east (Virginia) region, which I figure is about as close to NCBI as I can get. 
I've had three fastqdump processes running in parallel now for approximately 18 hours.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
